 CASARSAGURU/ISTOCK/GETTY IMAGES PLUS
CASARSAGURU/ISTOCK/GETTY IMAGES PLUS
Use of Proteomics in Dentistry
Human proteome mapping offers a sophisticated and highly individualized tool for predicting, diagnosing and managing oral and systemic diseases.
This course was published in the August 2021 issue and expires August 2024. The author has no commercial conflicts of interest to disclose. This 2 credit hour self-study activity is electronically mediated.
EDUCATIONAL OBJECTIVES
After reading this course, the participant should be able to:
- Describe the role of proteomics in disease detection, prevention, diagnosis and monitoring.
- Discuss applications for proteomics in dentistry.
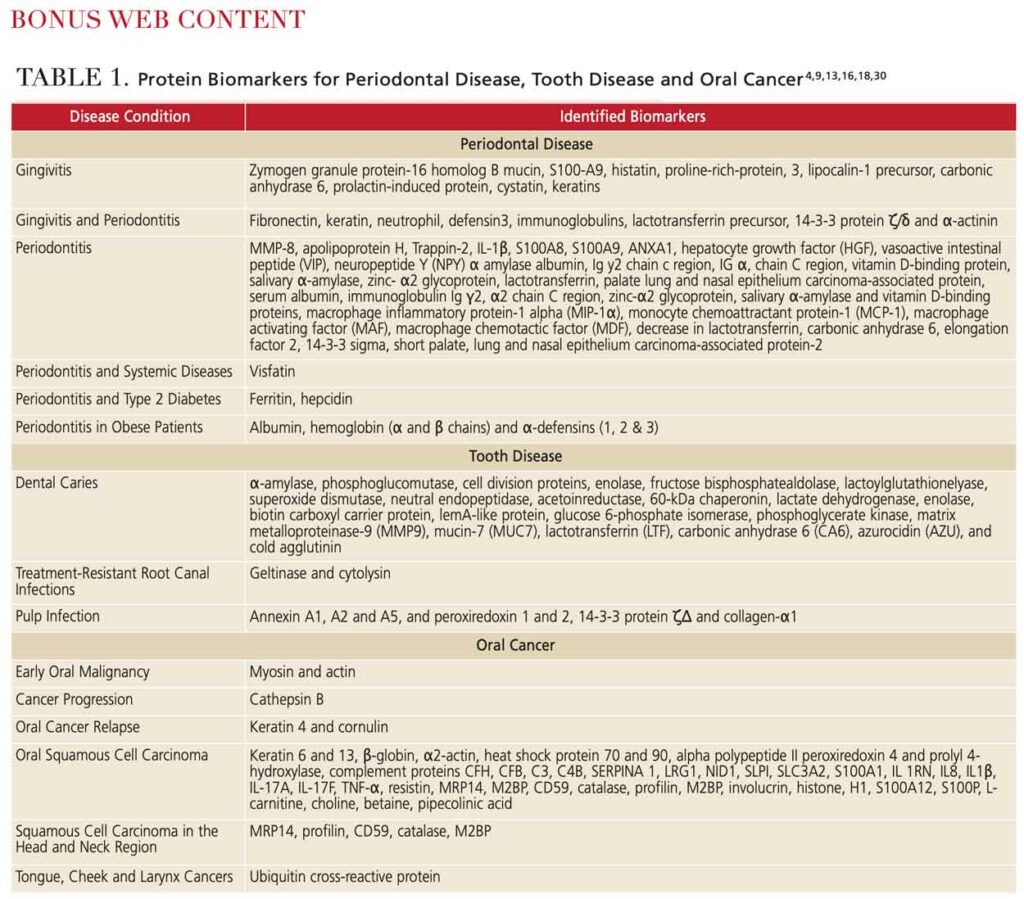
- Identify the protein biomarkers associated with oral diseases.
While genomic information has improved scientific knowledge of disease pathogenesis, genes do not fully reflect a cell’s status. The genome is essentially the operating manual that issues instructions for cells to make the specific proteins needed to carry out bodily functions.1 As such, proteins provide essential functional information and our “proteome identity.”2 Paradoxically, proteins can be the cause of disease (known as biomarkers), or used to cure disease (e.g., the use of antibodies as therapeutics). Proteins — such as hormones, antibodies, enzymes and cytokines — secreted in saliva by pathological cells serve as biomarkers for noninvasive disease screening. Thus, the study of protein structure, function and interaction, known as proteomics, is crucial for early detection and improved diagnosis, prognosis and monitoring of diseases. This discipline also contributes to the development of precise, highly effective therapies.
In the future, diagnosis of most diseases will not come from measuring the genetic blueprint, but, rather, from measuring proteins in their original and modified forms, as well as studying the interactions that determine cell and tissue status or disease stage. The ability of proteomics and genomics to examine human disease on a molecular level and determine individual response to disease and therapy has opened the door for a new approach to therapeutics and disease treatment called precision medicine.2–5
According to Yoithapprabhunath et al,4 Beadle and Tatum suggested a link between genes, proteins and disease in 1941; however, Wilkins coined the term proteome — which blends the words protein and genome — in 1994. In 2020, the Human Proteome Organization (HUPO) marked the 10th anniversary of the Human Proteome Project by reporting the identification of 17,874 proteins, representing 90% of the predicted 19,773 proteins coded in the human genome. Of the proteins identified so far, only 1254 do not have a known function. The missing 10% represent 1899 proteins without sufficient evidence of their existence.1 The goal of HUPO is twofold. First, it aims to map all human proteins present in health and disease, and, second, to determine all functions and interactions in their original and modified forms.
Identifying the human proteome is more complicated than identifying the human genome because protein structure is more complex. It is subject to chemical modifications, and may only be expressed at certain times or under certain conditions. In addition, not all proteins are found in (nearly) every human cell, such as with DNA.1 In-depth knowledge of the proteome will allow the comparison of proteins expressed by one person to those of another and under different conditions and disease states. Tests used to detect pregnancy, high cholesterol, strep throat, influenza, human immunodeficiency virus or prostate cancer are examples of medical assays that look for the presence of a single protein.
In the future, an individual’s proteome will be analyzed to identify abnormalities and treat them by modifying proteins and allowing the body to heal itself.6 With more than 99.9% of DNA being shared by every human, proteomics holds the key to answering how 0.1% of genes in each individual controls the countless phenotypes, from eye color to risk for developing disease.4,5 Already, the study of the genome and proteome has led to the development of new drugs through the identification of protein biomarkers associated with disease, and then designing agents to interfere with their actions.7 Molecular corrector drugs, such as elexacaftor/tezacaftor/ivacaftor (which were launched in late 2019 for treating cystic fibrosis), demonstrate how a drug can correct a defective amino acid in a protein chain and restore normal protein function.8

PROTEOMICS IN DENTISTRY
Precision medicine promotes customized healthcare via decisions and treatments tailored to the needs of the individual. To implement precision medicine in dentistry, clinicians must be knowledgeable about genomics, proteomics, and the relationship between oral and systemic health, and also be proficient in the use of new diagnostic tools. Analysis of a patient’s genome, proteome, environmental factors and cultural influences is used to develop personalized therapy, prescribe more rationally, dose more appropriately, and predict patient response.9
Researchers are looking to map proteins expressed in teeth and dental structures to gain an in-depth understanding of how the proteome can be used to monitor health status, disease onset, treatment response and outcomes. Once known, a complete list of proteins for the oral cavity and their functions will include thousands of entries.4,5 To date, the primary focus of dental proteomic research has been identification of proteins and their differential expression between disease and health.10 Although many of the identified biomarkers are awaiting further validation and have not been adopted in the clinical setting, further research and commercialization are likely to revolutionize dentistry by making early diagnosis predictably routine. Traditional biomarkers used to identify oral disease will soon be replaced by those enabling mitigation of disease and administration of targeted therapeutics.4,9
Proteomic research has identified more than 1000 salivary protein biomarkers, including secretory immunoglobulin A, lactoperoxidase, statherin, proline-rich glycoprotein, truncated cystatin S, cystatins, lysozyme and histatin-5. Saliva proteomics allows convenient monitoring of the patient’s physiology and prognostic and diagnostic biomarkers that may improve quality of life and reduce the financial burden associated with chronic disease. In addition to salivary testing, the oral fluid nanosensor test (OFNASET), or lab on a chip, is a handheld device capable of detecting up to eight salivary biomarkers in less than 15 minutes.11–14 Research indicates this technology effectively identifies biomarkers, but additional evidence is needed to confirm sensitivity, specificity, reproducibility, and correlation with existing disease diagnostic criteria. Although the intended use of OFNASET is detection of salivary biomarkers for oral cancer, this technology shows promise in facilitating a wide range of in-office, real-time disease detection assays.14 Another emerging point-of-care technology is a test for periodontal disease that analyzes saliva for matrix metalloproteinase-8 (MMP-8) inflammation biomarkers in less than 60 seconds. The analysis report offers patients insight into treatment needs and provides assessment data corresponding to the 2018 American Academy of Periodontology grading classifications.15
Current challenges for proteomics in dentistry include identifying disease-specific biomarkers, establishing sensitivity and specificity of tests, and standardizing collection and storage of saliva samples —not to mention lack of approval by the U.S. Food and Drug Administration.16 Barriers to implementation of precision dentistry include inadequacies in reimbursement, lack of an integrated electronic health record, protection of genetic information, disparities in access by marginalized populations, and lack of patient interest.17,18 As precision dentistry progresses, public awareness and acceptance of personal genome sequencing and continued proteomic research focusing on prevention, treatment and prognosis are essential for success.18,19
CARIES PROTEOMICS
Proteins associated with enamel, cementum, pulp and dentin give insight into the regenerative ability of dental tissues. Research indicates that 72 proteins have been identified in enamel, 510 in cementum, 5097 in pulp, and 813 in dentin.20–23 Enamel-specific proteins are critical for proper enamel formation, yet proteomics is still determining the roles of the numerous proteins involved and the exact protein composition of healthy tooth enamel in all stages of development.24 Researchers propose 40% to 60% of susceptibility to caries is genetically determined.9 This includes defects in the following genes: amelogenin X-linked is responsible for production of amelogenin, which ensures normal enamel development; kallikrein-related peptidase 4 (KLK4) is involved in enamel maturation; lysozyme-like protein 2, which produces lysozyme-2 protein involved in antibacterial defenses; and adherens junctions associated protein 1, which is associated with tooth development.9 Major enamel proteins that have been identified include amelogenin, ameloblastin, enamelin and tuftelin. Additional proteins associated with enamel formation include endoplasmic reticulum protein 29, calbindin, MMP-20 and KLK4.12,24 The identification of proteins exclusive to caries-free versus caries-prone individuals may aid in understanding the mechanism of caries, risk evaluation, and advancing the science of caries prevention and management.
One proposed caries prevention strategy aims to block or weaken enzymes that enable pathogenic microbes to form biofilm or attach to the tooth surface.25 Early detection of biomarkers associated with caries gives oral health professionals the insight to diagnose and treat patients based on specific findings and, thus, better determine prognosis.19 Furthermore, caries risk factors can be used to stratify patients and personalize therapy.25 Researchers have found a mechanism, associated with the delta-like non-canonical notch ligand 1 gene, that offers a potential solution for tooth repair by enhancing stem cell activation and tissue regeneration. In the future, this mechanism may provide a means to repair teeth damaged by caries or trauma.26
Differences have been found between cementum proteomes of deciduous and permanent teeth. Of the 510 proteins identified in cementum, 123 are exclusive to deciduous teeth, including myeloperoxidase. In comparison, decorin and osteocalcin are two of the 128 proteins exclusive to permanent teeth, and an additional 259 are expressed by both. These findings provide a greater understanding of differences between deciduous and permanent cementum, as well as insight into physiological/pathological events, such as root resorption.21
The dentin and pulp function as a complex, with odontoblasts from peripheral pulp tissue extending into the dentin, which is a unique connective tissue with specific proteins, such as dentin sialophosphoprotein, as well as proteins also found in bone, such as glycoprotein and osteonectin. Proteins mapped in dentin include dentin sialophosphoprotein, biglycan, osteopontin and osteocalcin. Their functions include cytoskeletal protein binding, immune transport, calcium and ion binding, and formation of extracellular matrix.22 Proteins mapped in dental pulp vary between health, inflammation and necrosis. An inflammatory process occurs when pulp disease advances, as evidenced by an abundance of immunoglobulins, including annexin A1, A2 and A5, and peroxiredoxin 1 and 2. Concomitantly, a repair process triggers upregulation of 14-3-3 protein ζΔ and collagen-α1 that may be associated with platelet activation and odontogenesis.27
Mapping the proteome dentin-pulp complex will aid in scientific understanding of its regenerative potential in response to disease and trauma, identify the stages of pathology, and help define biomarkers for new regenerative therapies in endodontics.22,27
PERIODONTAL PROTEOMICS
The one-size-fits-all approach of traditional assessment and nonsurgical periodontal therapy may not adequately treat those at greatest risk for periodontal disease — and may overtreat those at low risk for periodontal disease progression. Proteomics aims to identify specific biomarkers that enable decision-making based on hard data, rather than subjective assessments.
Current research shows genetic factors play a role in the clinical variability of periodontal disease by affecting host-bacterial interactions.9 Considered a polygenic disease, periodontitis is linked to a number of proteins.13 Research is focused on the proinflammatory cytokine interleukin-1 (IL-1) and MMP-8.9,15,28 Approximately 30% of the population is positive for the IL-1 genetic marker.29 Individuals who carry the IL-1 gene experience higher incidence and increased severity of periodontal disease.9 Currently, MMP-8 is considered the most promising biomarker for early diagnosis of periodontitis and peri-implantitis.15 A recent study showed that MMP-8 had a stronger correlation to bleeding on probing than plaque levels, thereby making it less susceptible to confounding effects of oral hygiene.28
Alveolar bone, with an identified 2625 proteomes, is critical for tooth retention. Currently, bone grafts are commonly used for alveolar bone regeneration therapy. Therefore, developing techniques that do not require surgery is critical. Transfer of the bone morphogenetic protein‐2/7 gene into periodontal tissues may cause stem cells in periodontal tissue to differentiate into osteogenic cells, enabling generation of new alveolar bone.30,31 This represents a promising nonsurgical approach.
Over time, the etiology of periodontitis has moved from biofilm to host-bacterial interaction and host-immune response. The next shift will involve proteomics and precision medicine. Proteomics offers hope that someday periodontitis risk can be determined early so disease can be prevented. At present, tests are available to screen for IL-1 and MMP-8. By integrating the precision medicine model into preventive care, time and resources can be allocated efficiently to patients at highest risk.9,13
ORAL CANCER PROTEOMICS
Developments in this field are helping clinicians predict which oral lesions are likely to become malignant. A number of protein biomarkers for oral cancer have been studied, including, but not limited to, complement proteins and inflammatory cytokines.13,32–34 The direct contact between saliva and oral cancer lesions allows for detection of biomarkers.3 In head and neck cancer, specific proteins show promise in predicting the biomarkers used to determine drug targets and therapeutic response, as well as in assessing risk for cancer recurrence.9 A prime example of how proteomics is changing drug therapies is the biologic drug cetuximab, which is approved for treatment of oral squamous cell carcinoma. As a monoclonal antibody, cetuximab inhibits receptor binding of a protein responsible for tumor cell growth.35
CONCLUSION
As evidenced by recent developments in the field, application of proteomics will soon become commonplace. Research suggests proteomics can be a means to predict, diagnose and monitor a number of oral and systemic diseases. This approach aims to augment or replace traditional clinical and radiographic assessments with low sensitivity and low positive predictive values that only assess history, extent and severity of disease. Precision medicine shows promise in diagnosing and managing oral diseases, as well as systemic conditions that affect the oral cavity.
Pinpointing the exact biomarkers associated with various diseases and ensuring high sensitivity and specificity are vital to application of proteomics in a clinical setting. In the near future, point-of-care salivary testing or other methodologies will assess the patient’s proteome identity quickly, aiding oral health professionals in diagnosis, treatment planning and delivery of optimal care.
REFERENCES
- Overall C. The human proteome: 90% in the light, 10% on the dark side. J Proteome Res. 2020;19:4731−4734.
- Alaaeddine R, Favad M, Nehme E, et al. The emerging role of proteomics in precision medicine: applications in neurodegenerative diseases and neurotrauma. Adv Exp Med Biol. 2017;1007:59–70.
- Li Q, Ouyang X, Chen J, et al. A review on salivary proteomics for oral cancer screening. Curr Issues Mol Biol. 2020;37:47–56.
- Yoithapprabhunath TR, Nirmal NM, Santhadevy A, et al. Role of proteomics in physiologic and pathologic conditions of dentistry: overview. J Pharm Bioallied Sci. 2015;7(Suppl 2):S344–S349.
- Overall C. The HUPO high stringency inventory of humanity’s shared human proteome revealed. J Proteome Res. 2020;19:4211–4314.
- Wilson TG Jr. Proteomics is reshaping health care. Decisions in Dentistry. 2019;5(4):6.
- National Human Genome Research Institute. Introduction to Genomics. Available at: genome.gov/About-Genomics/Introduction-to-Genomics#:~:text=What’s%20a%20Genome%3F,that%20organism%20throughout%20its%20life. Accessed June 29, 2021.
- DeWeerdt S. Cystic fibrosis drugs target the malformed proteins at the root of the disease. Nature. 2020;583:S2–S4.
- Reddy MS, Shetty SR, Vannala V, et al. Embracing personalized medicine in dentistry. J Pharm Bioall Sci. 2019;11:S92–S96.
- Rizal MI, Soeroso Y, Sulijaya B, et al. Proteomics approach for biomarkers and diagnosis of periodontitis: systematic review. Heliyon. 2020;6:1–8.
- Kaczor-Urbanowicz KE, Carrera-Presas CM, Aro K, et al. Saliva diagnostics — current views and direction. Exp Biol Med. 2017;242:459–472.
- Kaczor-Urbanowicz KE, Carrera-Presas CM, Kaczor T, et al. Emerging technologies for salivaomics in cancer detection. J Cell Mol Med. 2017;21:640–647.
- Katsani KR, Sakellari D. Saliva proteomics updates in biomedicine. J Biol Res-Thessaloniki. 2019;26:17.
- Daniel GS, Thiruppathy M, Aswath N, et al. Lab on a chip: Conquer disease at the earliest. J Pharm Bioall Sci. 2018;10:106–108.
- Alassiri S, Parnanen P, Rathnayake N, et al. The ability of quantitative, specific, and sensitive point-of-care/chair-side oral fluid Immunotests for aMMP-8 to detect periodontal and peri-implant diseases. Dis Markers. 2018;2018:1306396.
- American Dental Association. Salivary Diagnostics. Available at: ada.org/en/member-center/oral-health-topics/salivary-diagnostics. Accessed June 29, 2021.
- Polverini P. Personalized medicine and the future of dental practice. Per Med. 2018;15:449–451.
- Khurshid Z, Zohaib S, Najeeb S, et al. Advances in proteomic sciences in dentistry. Int J Mol Sci. 2016;17:728.
- Baheti A, Kambali S, Hattarki S, et al. Personalized medicine in dentistry. J Orofac Sci. 2015;9:3–6.
- Green DR, Schulte F, Lee KH, et al. Mapping the tooth enamel proteome and amelogenin phosphorylation onto mineralizing porcine tooth crowns. Front Physiol. 2019;10:925.
- Giovani PA, Martins L, Salmon CR, et al. Comparative proteomic analysis of dental cementum from deciduous and permanent teeth. J Periodont Res. 2021;56:173–185.
- Abbey SR, Eckhard U, Solis N, et al. The human odontoblast cell layer and dental pulp proteomes and N-terminomes. J Dent Res. 2018;97:338–346.
- Widbiller M, Schweikl H, Bruckmann A, et al. Shotgun proteomics of human dentin with different prefractionation methods. Sci Rep. 2019;9:1–8.
- Gil-Bona A, Bidlack FB. Tooth enamel and its dynamic protein matrix. Int J Mol Sci. 2020;21:1–25.
- Guedes SF, Neves BG, Bezerra DS, et al. Saliva proteomics from children with caries at different severity stages. Oral Dis. 2020;26:1219–1229.
- Walker JV, Zhuang H, Singer D, et al. Transit amplifying cells coordinate mouse incisor mesenchymal stem cell activation. Nat Commun. 2019;10:3596.
- Silva PA, Lima SM, Freire MS, Murad AM, Franco OL, Rezende TM. Proteomic analysis of human dental pulp in different clinical diagnosis. Clin Oral Investig. 2021;25:3285–3295.
- Sorsa T, Alassiri S, Grigoriadis A, et al. Active MMP-8 (aMMP-8) as a grading and staging biomarker in the periodontitis classification. Diagnostics (Basel). 2020;10:61.
- Mihaela SS, Gianina I, Liliana P, et al. Risk predictors in periodontal disease. Rom J Oral Rehab. 2017;9:89–96.
- Bell PA, Solis N, Kizhakkedathu JN, et al. Proteomic and N-terminomic tails analyses of human alveolar bone proteins: Improved protein extraction methodology and lysarginase digestion strategies increase proteome coverage and missing protein identification. J Proteome Res. 2019;18:4167–4179.
- Kawai M, Kataoka Y-H, Sonobe J, et al. Non‐surgical model for alveolar bone regeneration by bone morphogenetic protein‐2/7 gene therapy. J Periodontol. 2018;89:85–92.
- The Human Protein Atlas. The Head and Neck Cancer Proteome. Available at: proteinatlas.org/humanproteome/pathology/head+and+neck+cancer. Accessed June 29, 2021.
- Chu HW, Chang KP, Hsu CW, et al. Identification of salivary biomarkers for oral cancer detection with untargeted and targeted quantitative proteomics approaches. Mol Cell Proteomics. 2019;18:1796–1806.
- Singh P, Verma JK, Singh JK. Validation of salivary markers, IL-1β, IL-8 and Lgals3bp for detection of oral squamous cell carcinoma in an Indian population. Sci Rep. 2020;10:1–9.
- Taberna M, Oliva M, Mesia R. Cetuximab-containing combinations in locally advanced and recurrent or metastatic head and neck squamous cell carcinoma. Front Oncol. 2019;9:1–13.
From Decisions in Dentistry. August 2021;7(8)26-28, 31.




